Cell Free DNA Analysis
While at UAB I designed a magnetic bead pull down assay utilizing peptide nucleic acids that could isolate and purify telomeric DNA based on either the quadruplex structure or on primary sequence. This assay is the first of its kind due to its flexibility to isolate any quadruplex forming DNA from the cell free matrix based on structural recognition or by targeting the sequence to isolate a specific quadruplex forming DNA fragment such as the telomere sequence. The general bead capture scheme can be found in figure 1.
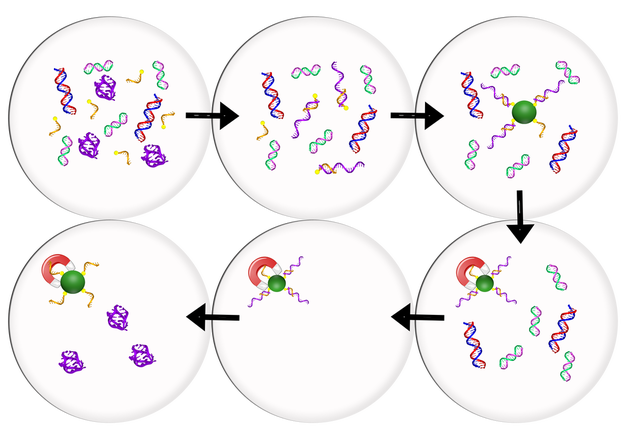
Figure 1. The generalized procedure for DNA capture and purification with magnetic beads. In this method a biotinated capture strand complementary to our target strand is added under denaturing conditions. Denaturant is removed allowing the target strand to hybridize with the capture strand. Streptavidin beads are then added, allowing the biotin tag to bind to the magnetic bead, thus isolating our target strand. Once a magnetic field is applied to the sample, the beads are fixed to the vial wall and the remaining impurities can be removed and rinsed from the sample. The now pure and isolate sequence can then be released from the bead by adding denaturant to de-hybridized the target strand from the biotinated capture stand.
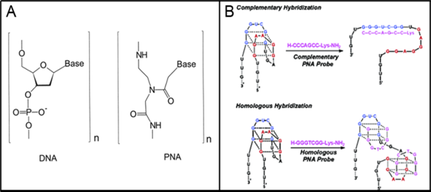
This assay has novel flexibility because of the use of peptide nucleic acids developed by our collaborator Dr. Bruce Armitage, Professor of Chemistry at Carnegie Mellon University in Pittsburgh, PA. The Armitage laboratory is a world leader in the design and synthesis of peptide nucleic acids (PNAs), nucleic acid like molecules that can hybridize to target DNA and RNA molecules. As shown in Figure 2A, instead of the normal sugar-phosphate backbone as found in native nucleic acids, PNAs contain a peptide-like backbone. The bases attached to each PNA unit are oriented in such that they can form regular Watson-Crick base pairs with either complementary DNAs or RNAs with high selectivity and affinity while also imparting nuclease and protease resistance. Armitage and coworkers have also demonstrated that PNAs can be designed to hybridize with G-quadruplex DNA either through Watson-Crick base pairing with a complementary strand to form a DNA/PNA duplex or alternatively through the formation of Hoogstein base pairs with a PNA G-quadruplex forming a hybrid PNA/DNA G-quadruplex as shown in Figure 2B.
Preliminary findings from my graduate work demonstrate that utilizing PNAs to pull down quadruplex DNA and telomeric DNA specifically is effective at ranges of 70-80% capture efficiency with a release of approximately 90%. However, I would like to see this assay increase to above 95% capture efficiency and 99% release efficiency so that we could utilize this technique for not only the isolation and purification of quadruplex and telomeric DNA in cell-free or serum samples but also eventually as a quantitative tool to measure telomeric DNA in any sample matrix. In order to do this, I have continued collaborations with Dr. Armitage to developed γ-modified PNA ligands which enchances to PNA/DNA binding affinity to fentamolar concentrations. We plan to explore various PNA sequences and structural motifs to enhance binding and release, as well as “catch and release” motifs where bound DNA can be released from the PNA by adding a fully complementary PNA strand that will bind to the bead bound PNA, displacing the capture DNA.
This bead capture assay development project requires minimal instrumentation, only needing a UV-Vis spectrophotometer to monitor DNA capture and release, and possibly a fluorometer to monitor fluorescently labeled PNA assays, and can be done on campus. Just as in the quadruplex characterization project, we would also like to further characterize the PNA/DNA interactions which will expand the scope of this project and expose students to instrumental techniques described previously and can be done with the instrumentation available to me at UAB. Eventually we could expand this project further by creating PNA capture strands for other known DNA sequences of interest such as known genetic mutations or cell-free fragments that could be used as biomarkers in serum samples. This research project will give my students valuable hands on experience with a wide range of instrumentation and general lab technique and can also be easily modified to meet the work demands of each individual student who is interested in participating.
This bead capture assay development project requires minimal instrumentation, only needing a UV-Vis spectrophotometer to monitor DNA capture and release, and possibly a fluorometer to monitor fluorescently labeled PNA assays, and can be done on campus. Just as in the quadruplex characterization project, we would also like to further characterize the PNA/DNA interactions which will expand the scope of this project and expose students to instrumental techniques described previously and can be done with the instrumentation available to me at UAB. Eventually we could expand this project further by creating PNA capture strands for other known DNA sequences of interest such as known genetic mutations or cell-free fragments that could be used as biomarkers in serum samples. This research project will give my students valuable hands on experience with a wide range of instrumentation and general lab technique and can also be easily modified to meet the work demands of each individual student who is interested in participating.